Mummichog Server
pathway and network analysis for metabolomics
Updating cloud backend... please excuse downtime.
One may use our v3 test server at the meantime.
Start new analysis
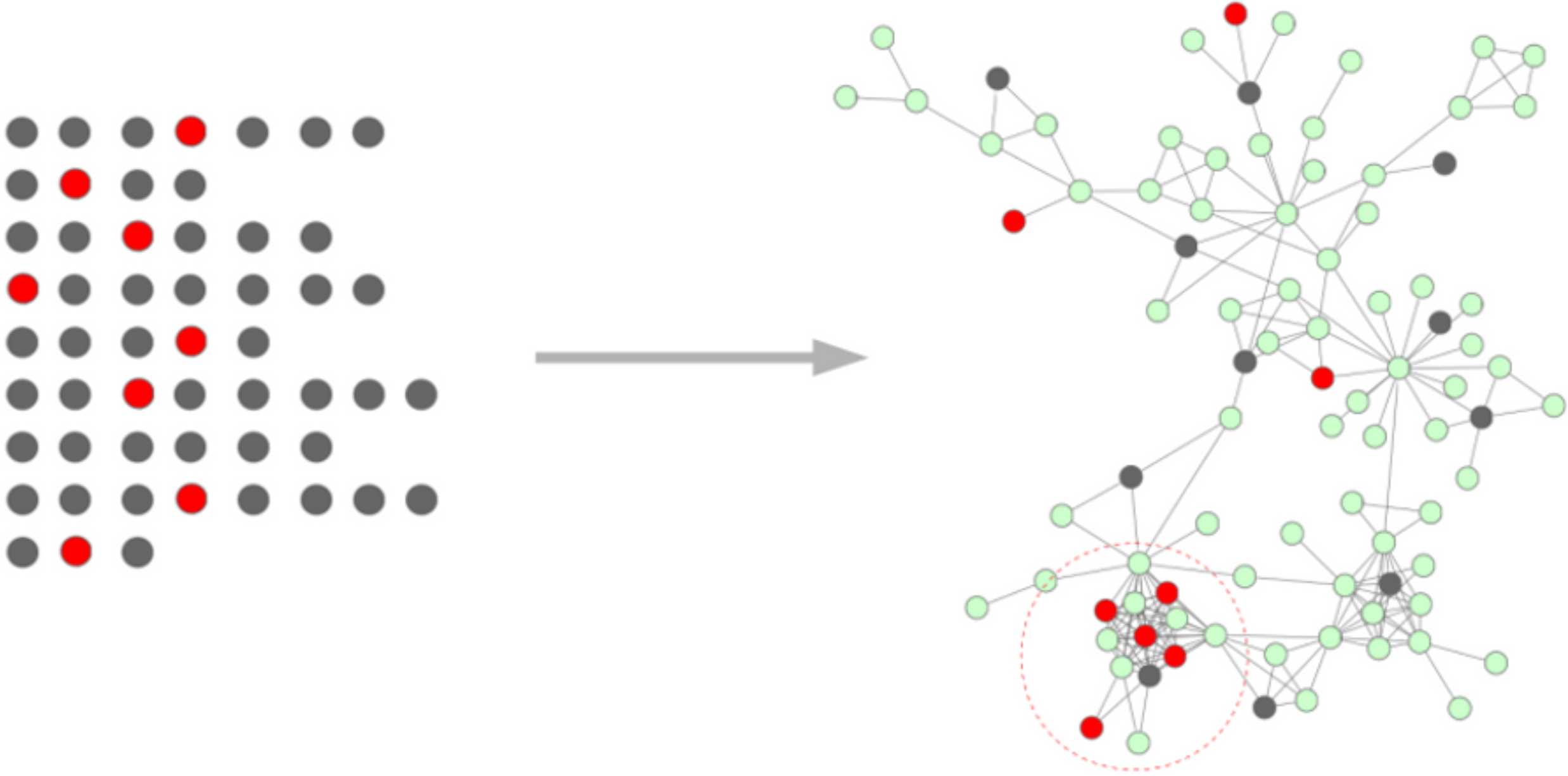
Mummichog searches
for enrichment patterns on metabolic network,
bypassing metabolite identification, to generate high-quality hypotheses directly from a LC-MS data table.
Format of input file (right click to save an example):
| m/z | retention time | p-value | statistic |
|---|---|---|---|
| 85.0278 | 59 | 0.002657 | -3.55 |
| 85.0472 | 124 | 0.730810 | -0.35 |
| 85.0653 | 68 | 0.086509 | 1.83 |
| 85.1007 | 16 | 0.057916 | -2.04 |
| 86.0595 | 67 | 0.076789 | -1.89 |
Please cite this paper if you use mummichog: Li et al. (2013) Predicting network activity from high throughput metabolomics. PLoS computational biology 9.7 (2013): e1003123.
The author can be reached at shuzhao.li at Gmail.